Chapter 8 of 10 - AP Biology Course
Gene Expression & Regulation
Every cell in your body carries the same DNA, yet a neuron looks and functions nothing like a liver cell. This chapter traces the central dogma from DNA structure and replication through transcription and translation, then examines how prokaryotic and eukaryotic cells regulate which genes are expressed, when, and how much. It concludes with biotechnology tools that exploit these processes.
DNA Structure
DNA is a double-stranded helix with an antiparallel orientation: one strand runs 5' to 3' while the complementary strand runs 3' to 5'. The backbone consists of alternating deoxyribose sugars and phosphate groups linked by phosphodiester bonds. Nitrogenous bases project inward and pair via hydrogen bonds: adenine (A) with thymine (T) using two hydrogen bonds, and guanine (G) with cytosine (C) using three.
This complementary base pairing is the foundation for accurate replication, transcription, and every hybridization-based technology (PCR primers, DNA probes, CRISPR guide RNA). The antiparallel arrangement means enzymes like DNA polymerase can only synthesize in the 5' to 3' direction, which creates the leading-strand and lagging-strand distinction during replication.
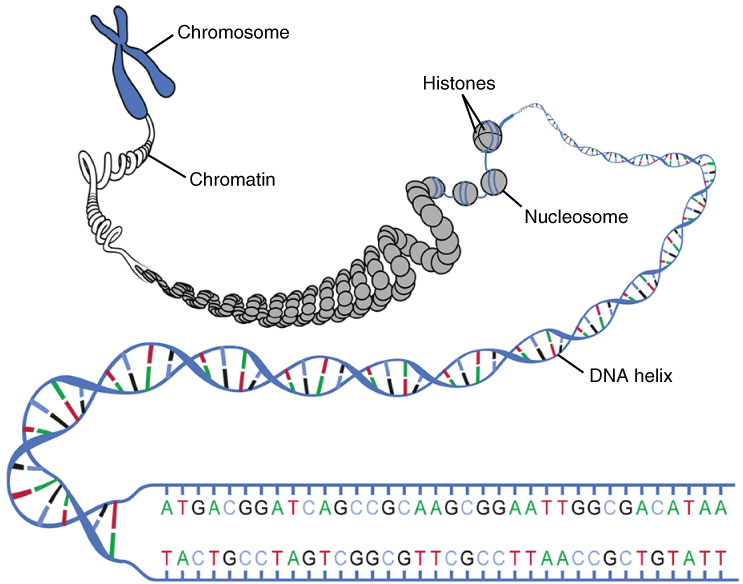
DNA macrostructure: the double helix is wound around histone proteins to form nucleosomes, which compact further into chromatin and ultimately into visible chromosomes during cell division.
DNA Replication
Replication is semiconservative: each daughter molecule retains one original parent strand paired with one newly synthesized strand, as demonstrated by the Meselson-Stahl experiment. The process begins at origins of replication, where helicase unwinds the helix and single-strand binding proteins stabilize the separated strands.
DNA polymerase III adds nucleotides in the 5' to 3' direction. The leading strand is synthesized continuously toward the replication fork. The lagging strand is synthesized in short segments called Okazaki fragments, each initiated by an RNA primer laid down by primase. DNA polymerase I replaces primers with DNA, and DNA ligase seals the nicks between fragments.
Telomeres are repetitive DNA sequences at chromosome ends that shorten with each round of replication because the lagging strand cannot be fully replicated at the terminus. The enzyme telomerase extends telomeres in germ cells and stem cells, preventing critical shortening. Most somatic cells lack significant telomerase activity, which contributes to cellular aging.
Transcription
During transcription, RNA polymerase binds the promoter region upstream of a gene, unwinds a short stretch of DNA, and synthesizes a complementary RNA strand in the 5' to 3' direction using the template (antisense) strand. Transcription continues until RNA polymerase reaches a terminator sequence.
In eukaryotes, the primary transcript (pre-mRNA) undergoes three major modifications before leaving the nucleus: a 5' methylguanosine cap is added to protect against degradation and aid ribosome binding, a poly-A tail (100-250 adenine nucleotides) is added to the 3' end for stability, and introns are spliced out by the spliceosome while exons are joined. Alternative splicing allows one gene to encode multiple protein variants.
Translation
Translation converts mRNA codons into a polypeptide chain at the ribosome. Each codon (three-nucleotide sequence) specifies an amino acid or a stop signal. Transfer RNA (tRNA) molecules carry amino acids and have anticodons that base-pair with mRNA codons.
Translation begins when the small ribosomal subunit binds the mRNA and locates the start codon (AUG), which codes for methionine. The large subunit joins, and elongation proceeds: tRNAs enter the A site, peptide bonds form, the ribosome translocates, and the growing chain emerges. Translation ends when a stop codon (UAA, UAG, or UGA) enters the A site and release factors trigger polypeptide release.
The flow of genetic information in cells. Each arrow represents a distinct molecular process catalyzed by specific enzymes.
DNA
Double-stranded template storing genetic information in base sequence.
Transcription
RNA polymerase copies the template strand into a complementary mRNA.
mRNA processing
5' cap, poly-A tail, and intron splicing (eukaryotes only).
Translation
Ribosomes read codons; tRNAs deliver amino acids; polypeptide forms.
Protein
Folded polypeptide performs structural, enzymatic, or regulatory functions.
Mutations
Mutations are permanent changes in the DNA sequence. They range from single-nucleotide substitutions to large chromosomal rearrangements. At the gene level, the AP exam focuses on point mutations (single base changes) and frameshift mutations (insertions or deletions that shift the reading frame).
| Mutation type | What changes | Effect on protein | Example |
|---|---|---|---|
| Silent | One base substituted | No amino acid change (codon redundancy) | GCU to GCC both code for alanine |
| Missense | One base substituted | Different amino acid incorporated | Sickle cell: GAG to GUG (Glu to Val) |
| Nonsense | One base substituted | Premature stop codon, truncated protein | UAC (Tyr) to UAA (stop) |
| Frameshift | Insertion or deletion (not multiple of 3) | All downstream codons altered | Cystic fibrosis delta-F508 deletion |
Gene Regulation in Prokaryotes
Bacteria regulate transcription primarily at the operon level. An operon is a cluster of genes under the control of a single promoter and operator. The lac operon is an inducible system: when lactose is absent, the repressor binds the operator and blocks transcription. When lactose is present, its derivative allolactose binds the repressor, causing a conformational change that releases it from the operator, allowing RNA polymerase to transcribe the structural genes (lacZ, lacY, lacA).
The trp operon is a repressible system. When tryptophan is abundant, it acts as a corepressor by binding the trp repressor protein, activating it to bind the operator and shut down transcription. When tryptophan levels drop, the repressor cannot bind the operator, and the biosynthetic enzymes are produced.
Eukaryotic Gene Regulation
Eukaryotic regulation is more complex and occurs at multiple levels. Enhancers are distant DNA sequences where activator proteins bind to increase transcription rate. Silencers bind repressor proteins that decrease transcription. Mediator proteins and transcription factors assemble at the promoter as part of a large initiation complex.
Epigenetic modifications alter gene expression without changing the DNA sequence. DNA methylation typically silences genes by preventing transcription factor binding. Histone acetylation loosens chromatin structure (euchromatin), making DNA more accessible. Histone deacetylation and methylation tighten chromatin (heterochromatin), reducing gene expression.
Post-transcriptionally, microRNA (miRNA) molecules bind complementary mRNA sequences and either degrade the mRNA or block its translation, providing another layer of expression control. RNA interference (RNAi) is a related mechanism used in research to silence specific genes experimentally.
Thymine (DNA base)
Thymine is one of the four nitrogenous bases in DNA. It pairs with adenine via two hydrogen bonds and is replaced by uracil in RNA - a key distinction between the two nucleic acids.
Formula
C5H6N2O2
Mol. Weight
126.11 g/mol
Biotechnology Tools
PCR (Polymerase Chain Reaction) amplifies a specific DNA segment in vitro using heat-stable DNA polymerase (Taq), primers flanking the target region, and repeated cycles of denaturation (95 C), annealing (50-65 C), and extension (72 C). Each cycle doubles the target, producing millions of copies within hours.
Gel electrophoresis separates DNA fragments by size. Smaller fragments migrate farther through the agarose matrix toward the positive electrode (DNA is negatively charged). This technique is used alongside restriction enzymes to analyze DNA, create genetic fingerprints, and verify PCR products.
CRISPR-Cas9 is a genome-editing tool derived from bacterial immune systems. A guide RNA directs the Cas9 nuclease to a complementary DNA sequence, where it creates a double-strand break. The cell's repair machinery then introduces insertions, deletions, or precise edits. CRISPR has transformed genetics research and holds therapeutic potential for correcting disease-causing mutations.
Quick Check
Which strand of DNA is used as the template during transcription?
Fill in the Blank
On the lagging strand, short DNA segments synthesized in the 5' to 3' direction are called________. They are later joined by DNA ligase into a continuous strand.
Quick Check
A single base insertion in the middle of a gene would most likely cause which type of mutation?
Quick Check
In the lac operon, what molecule binds the repressor and prevents it from blocking transcription?
Fill in the Blank
Adding methyl groups to DNA typically________gene expression by preventing transcription factor access to the promoter.
Was this helpful? Rate it!
Create a free Lorea account
Turn your notes into courses, practice tests, study games, and narrated videos - or build full interactive study worlds - then publish, download, and share them however you like.